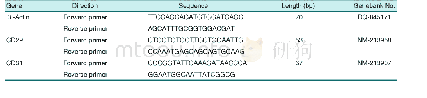
《Table 1 Genes and specific primers used for real-time polymerase chain reaction》
 提示:宽带有限、当前游客访问压缩模式
提示:宽带有限、当前游客访问压缩模式
本系列图表出处文件名:随高清版一同展现
《Identification of Genes Associated with Clubroot Resistance in Chinese Cabbage》
The total mapped reads were aligned to each region in the reference genome,including the exon,intergenic and intron regions,with the percentage of the exon region being the highest in all the four samples libraries(Fig.4).GC%of all the four sequences libraries was approximately 47%and Q30 percentage was above 88.52%(Table 2).A total of 13 405 116and 13 018 837 clean reads were obtained from SC-T and SC-CK,respectively;12 637 292 and 13 811 534clean reads were obtained from RC-T and RC-CK,respectively.The mapped ratios of these four samples to the reference database for B.rapa genome were all above 80%(Table 2).These results suggested that the sequence data were reliability.
| 图表编号 | XD0014158700 严禁用于非法目的 |
|---|---|
| 绘制时间 | 2018.12.25 |
| 作者 | Song Bo、Xu Hai、He Jiang-ming、Hu Jing-feng、Fan Xiao-xue、Chen Long-zheng |
| 绘制单位 | Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences、Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences、Institute of Horticultura Crops, Yunnan Academy of Agricultural Sciences、Institute of Horticultura Crops, Yunnan |
| 更多格式 | 高清、无水印(增值服务) |
查看“Table 1 Genes and specific primers used for real-time polymerase chain reaction”的人还看了
-
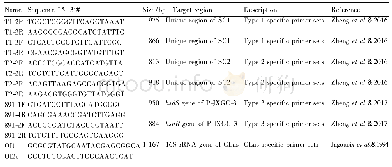
- Table 1 General information ofthe prophage-type specific primers and16S r DNA-specific primers used in this study