《Table 3 Number of differentially expressed genes detected in sequence analysis》
 提示:宽带有限、当前游客访问压缩模式
提示:宽带有限、当前游客访问压缩模式
本系列图表出处文件名:随高清版一同展现
《Identification of Genes Associated with Clubroot Resistance in Chinese Cabbage》
+shows gene expression changes detected;ND shows not detected.
Among 1 494 DEGs in SC upon inoculation,1 310were detected only in SC and 184 were shared with RC.Furthermore,734 of the 1 310 DEGs(56.0%)in SC were up-regulated,which was greater than the numbers of down-regulated DEGs(576;44.0%)(Table 3) .In RC,559 DEGs were unique to RC and184 were shared with SC.However,in contrast to SC,only 216 of 559 DEGs(38.6%)in RC were upregulated and 343 DEGs(61.4%)were down-regulated(Table 3),which suggested that fewer genes were upregulated in RC than in SC.These results suggested that RC might not have to up-regulate genes to resist pathogen infection.
| 图表编号 | XD0014158800 严禁用于非法目的 |
|---|---|
| 绘制时间 | 2018.12.25 |
| 作者 | Song Bo、Xu Hai、He Jiang-ming、Hu Jing-feng、Fan Xiao-xue、Chen Long-zheng |
| 绘制单位 | Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences、Institute of Vegetable Crops, Jiangsu Academy of Agricultural Sciences、Institute of Horticultura Crops, Yunnan Academy of Agricultural Sciences、Institute of Horticultura Crops, Yunnan |
| 更多格式 | 高清、无水印(增值服务) |
查看“Table 3 Number of differentially expressed genes detected in sequence analysis”的人还看了
-

- Table 1 Gene ontology enrichment analysis of methylation-regulated differentially expressed genes associated with colon
-
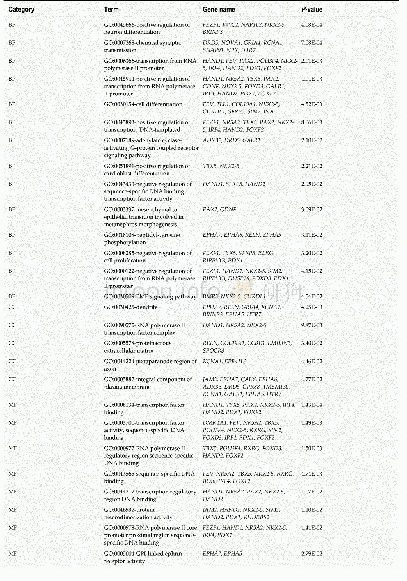
- Table 1 Gene ontology enrichment analysis of methylation-regulated differentially expressed genes associated with colon
-

- Table 2 Differentially expressed genes in the trans-regulatory networks existed in both GSE1420 and GSE26886 datasets





