《Table 1 Differentially expressed mi RNAs and target genes》
 提示:宽带有限、当前游客访问压缩模式
提示:宽带有限、当前游客访问压缩模式
本系列图表出处文件名:随高清版一同展现
《Differentially expressed miRNAs in premature infants with retinopathy-a bioinformatics analysis》
Data Source The data in our study came from the study of Wang et al[9]and Zhao et al[10].Wang et al[9]analyzed the mi RNA profile data of retinal tissue between 40 C57BL/6neonatal mice with ROP and 40 without ROP(controls)on postnatal day 17.Finally,67 differentially expressed mi RNAs were identified.Zhao et al[10]compared mi RNAs in neonatal Sprague-Dawley rats.They found 66 differentially expressed plasma mi RNAs by deep sequencing technology on postnatal day 14.Based on the above data,we perform a stricter screening standard which is the significant mi RNAs are selected from the intersection of the above two articles(Wang et al[9]and Zhao et al[10]).And fold change must fulfill greater than two in both studies.In addition,mi RNA sequence must be well conservative between human and rodent.After screening,only 1 up-regulated and 1 down-regulated mi RNAs passed the test,and were further selected into this study(Table 1).Based on these findings,we try to identify the roles of mi RNA in ROP.
| 图表编号 | XD0023059300 严禁用于非法目的 |
|---|---|
| 绘制时间 | 2018.05.18 |
| 作者 | Yang Yang、Jing-Jing Pan、Xiao-Guang Zhou、Xiao-Yu Zhou、Rui Cheng |
| 绘制单位 | Department of Neonatology,Children's Hospital Affiliated to Nanjing Medical University、Department of Neonatology,Jiangsu Provincial People's Hospital Affiliated to Nanjing Medical University、Department of Neonatology,Children's Hospital Affiliated to N |
| 更多格式 | 高清、无水印(增值服务) |
查看“Table 1 Differentially expressed mi RNAs and target genes”的人还看了
-

- Table 1 Gene ontology enrichment analysis of methylation-regulated differentially expressed genes associated with colon
-
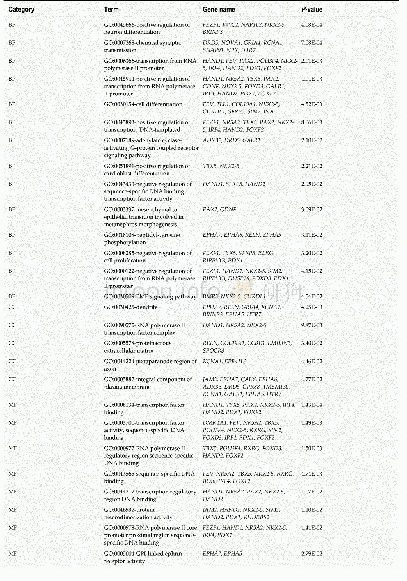
- Table 1 Gene ontology enrichment analysis of methylation-regulated differentially expressed genes associated with colon
-

- Table 2 Differentially expressed genes in the trans-regulatory networks existed in both GSE1420 and GSE26886 datasets





