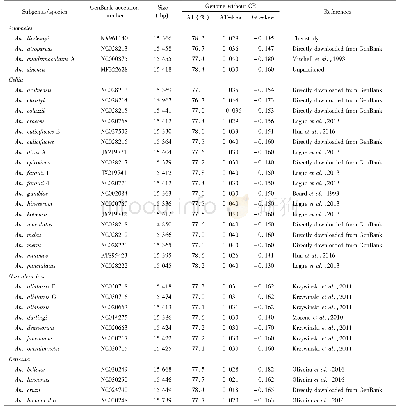
《Table 1–Soybean accessions used in the study.》
 提示:宽带有限、当前游客访问压缩模式
提示:宽带有限、当前游客访问压缩模式
本系列图表出处文件名:随高清版一同展现
《Development and validation of InDel markers for identification of QTL underlying flowering time in soybean》
Fifty six accessions,including 29 from three recent research papers and 27 from this study,were used for In Del polymorphism validation(Table 1).Young leaves from 27accessions were collected three weeks after planting in growth chambers and separately quick-frozen in liquid nitrogen.Total DNA was extracted by the improved cetyltrimethylammonium bromide(CTAB)method[36].A sequencing library was constructed with at least 6μg of genomic DNA following the manufacturer's instructions(Illumina Inc.,San Diego,CA).Paired-end sequencing libraries with an insert size of approximately 500 bp were sequenced on an Illumina Hi Seq 2000 sequencer.
| 图表编号 | XD0012315100 严禁用于非法目的 |
|---|---|
| 绘制时间 | 2018.04.01 |
| 作者 | Jialin Wang、Lingping Kong、Kanchao Yu、Fengge Zhang、Xinyi Shi、Yanping Wang、Haiyang Nan、Xiaohui Zhao、Sijia Lu、Dong Cao、Xiaoming Li、Chao Fang、Feifei Wang、Tong Su、Shichen Li、Xiaohui Yuan、Baohui Liu、Fanjiang Kong |
| 绘制单位 | The Key Laboratory of Soybean Molecular Design Breeding, Northeast Institute of Geography and Agroecology, Chinese Academy of Sciences、The Key Laboratory of Soybean Molecular Design Breeding, Northeast Institute of Geography and Agroecology, Chinese Acade |
| 更多格式 | 高清、无水印(增值服务) |



![Table 2 List of the Landsat data[22]used in this study](http://bookimg.mtoou.info/tubiao/gif/QQSJ201804005_29300.gif)