《Table 1.Development and analysis of T0 rice plants for targeted mutagenesis of Os LCT1 and OsNramp5
 提示:宽带有限、当前游客访问压缩模式
提示:宽带有限、当前游客访问压缩模式
本系列图表出处文件名:随高清版一同展现
《Characterization and Evaluation of OsLCT1 and OsN ramp5 Mutants Generated Through CRISPR/Cas9-Mediated Mutagenesis for Breeding Low Cd Rice》
a Due to technical reasons,we failed to amplify the fragment encompassing the target site in exon 9(Fig.1-B)for the other plants.
A total of 124 HPT-positive T0 plants were raised from transformation with the three CRISPR/Cas9 vectors(Table 1).Due to the difficulty in designing reliable primers for PCR amplification of the fragment encompassing the target site in exon 9 of Os Nramp5,a number of PCRs failed and consequently only four plants were included for mutation analysis of the 49T0 plants transformed with pH-nramp5×9(Table 1).Two-thirds of the mutated pH-lct1×1 T0 plants were monoallelic heterozygous and the remaining mutants carried biallelic heterozygous mutations,i.e.two different mutations at the same target site(Table 1).
| 图表编号 | XD0040097600 严禁用于非法目的 |
|---|---|
| 绘制时间 | 2019.03.28 |
| 作者 | LIU Songmei、JIANG Jie、LIU Yang、MENG Jun、XU Shouling、TAN Yuanyuan、LI Youfa、SHU Qingyao、HUANG Jianzhong |
| 绘制单位 | National Key Laboratory of Rice Biology、Institute of Nuclear Agricultural Sciences, Zhejiang University、Hubei Collaborative Innovation Center for Grain Industry、National Key Laboratory of Rice Biology、Institute of Nuclear Agricultural Sciences, Zhejiang U |
| 更多格式 | 高清、无水印(增值服务) |
查看“Table 1.Development and analysis of T0 rice plants for targeted mutagenesis of Os LCT1 and OsNramp5.”的人还看了
-
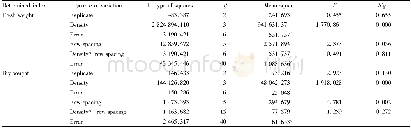
- Table 2 Variance analysis of the effects of different planting densities and row spacing on plant productivity (inter-su