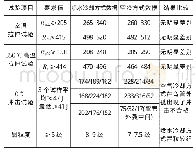
《Table 2 Comparison of properties of L(+)-dihydrobutanedioic acid-producing ORCHs from different spe
 提示:宽带有限、当前游客访问压缩模式
提示:宽带有限、当前游客访问压缩模式
本系列图表出处文件名:随高清版一同展现
《双头菌WH-1环氧乙烷二酸水解酶的克隆和酶学性质的研究(英文)》
Taken together,although the ORCH from Rhodococcus had the highest enzyme activity,its thermal and pH stability were the poorest.The properties of the ORCH from Klebsiella are intermediate between those of the enzymes from Labrys and Rhodococcus.The ORCH from Labrys sp.WH-1 showed the best catalytic efficiency and stability(thermal and pH stability),and is less sensitive to most metal ions and chemicals,indicating that it could be a good industrial biocatalyst for production of L(+)-dihydrobutanedioic acid.ORCH from Labrys sp.WH-1 was further cloned and its amino acid sequence was aligned with those from Rhodococcus and Klebsiella using ClustalW2 program(Fig.3).It showed 37%and 47%identity with the ORCHs from Rhodococcus and Klebsiella,respectively.The amino acid composition of these three ORCHs is shown in Table S2.The ORCH from Labrys has higher basic amino acid content(Arg,Lys,and His)than the other two,consistent with its higher isoelectric point(Fig.1d).The ORCH from Labrys sp.WH-1 has the highest hydrophobic amino acid content(Ala,Val,Ile,Leu,Phe,and Trp)and the lowest polar amino acid content(Cys,Ser,Thr,Tyr,Asn,and Gln)of these three enzymes,indicating that it has the best stability,which was confirmed by our experiments(the hydrophobic amino acid content is related to protein stability).Thesecondary structures of ORCHs were predicted by PredictProtein(http://www.predictprotein.org),and it gave similar results to those from the amino acid sequence alignment data(Fig.3),except for N-terminal sequences.UV CD was used to investigate the secondary structure,and the contents of helix and sheet in ORCH from Labrys sp.WH-1 were 40.3%and12.5%,respectively,which is consistent with software predictions.The nine residues for ORCH from Labrys sp.WH-1(D48,T52,R85,N165,K195,Y201,A219,H221,and D224)are completely conserved with the important catalytic residues for ORCHs from Rhodococcus(D18,T22,R55,N134,K164,Y170,A188,H190,and D193)and Klebsiella(D48,T52,R85,N165,K195,Y201,A219,H221,and D224),suggesting that they may adopt the same catalytic mechanism(Pan et al.,2011;Cheng et al.,2014b).
| 图表编号 | XD00113738000 严禁用于非法目的 |
|---|---|
| 绘制时间 | 2019.12.03 |
| 作者 | Wen-na BAO、Zi-sheng LUO、Shi-wang LIU、Yuan-feng WU、Pei-lian WEI、Gong-nian XIAO、Yong LIU |
| 绘制单位 | College of Biosystems Engineering and Food Science, Zhejiang University、School of Biological and Chemical Engineering, Zhejiang University of Science and Technology、Zhejiang Provincial Key Laboratory for Chemical and Biological Processing Technology of Fa |
| 更多格式 | 高清、无水印(增值服务) |